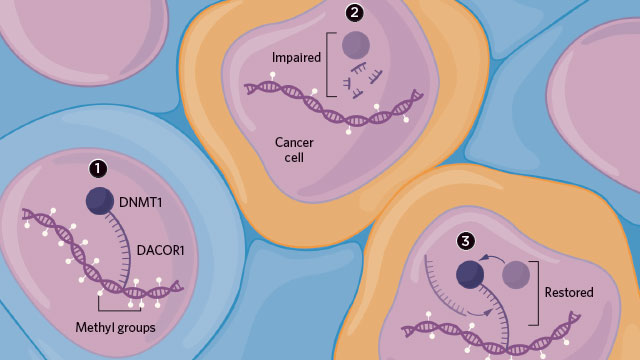
 UNSUPERVISED: In a healthy colon cell, the DNA methyltransferase DNMT1 interacts with the long noncoding RNA DACOR1 to properly methylate the cell’s genomic material (1). DACOR1 is downregulated in cancer cells, thus upsetting DNMT1’s activity and changing the cells’ methylation patterns to promote tumor growth (2). Adding DACOR1 back into cancer cells restores some normal methylation and gene expression (3).© LUCY READING-IKKANDA
UNSUPERVISED: In a healthy colon cell, the DNA methyltransferase DNMT1 interacts with the long noncoding RNA DACOR1 to properly methylate the cell’s genomic material (1). DACOR1 is downregulated in cancer cells, thus upsetting DNMT1’s activity and changing the cells’ methylation patterns to promote tumor growth (2). Adding DACOR1 back into cancer cells restores some normal methylation and gene expression (3).© LUCY READING-IKKANDA
EDITOR'S CHOICE IN GENETICS & GENOMICS
The paper
C.R. Merry et al., “DNMT1-associated long non-coding RNAs regulate global gene expression and DNA methylation in colon cancer,” Hum Mol Genet, 24:6240-53, 2015.
DNA methylation is a critical regulator of gene activity. In cancer cells, faulty methylation patterns contribute to the excessive growth and dysfunction characteristic of tumors. While increased methylation often occurs at sites of tumor suppressor genes—turning off their activity—reduced methylation is widespread throughout other parts of the genome, freeing up many sections of DNA to...
“No one understands what’s driving the hypomethylation” in cancer cells, says Ahmad Khalil, a geneticist at Case Western Reserve University in Ohio. “The initial hypothesis was that maybe the enzymes regulating histone modification or DNA methylation are mutated, or their expression is deregulated.” But many studies have found that mutations in these enzymes are rare, and their genes’ expression isn’t altered.
To determine how cancer cells undergo these massive changes in methylation, Khalil turned to long noncoding RNAs (lncRNAs). In a previous study, when Khalil’s group removed lncRNAs found in complexes with histone-modifying enzymes, genes that were normally repressed became upregulated. “I proposed the hypothesis that maybe the deregulation is of the lncRNAs, not [of] the enzymes that create these modifications,” he says.
Christopher Maher, a computational biologist at Washington University in Saint Louis who was not part of the study, says Khalil’s approach to look at how lncRNAs interact with DNA methyltransferases “makes a lot of sense biologically.”
There are three distinct DNA methyltransferase enzymes that attach a methyl group to cytosines in mammals; Khalil chose to examine one called DNMT1. With his colleagues, Khalil isolated DNMT1 and its associated RNAs from a colon cancer cell line and found 148 DNMT1-associated lncRNAs—but that was far too many to study individually. The researchers pored over data from the Cancer Genome Atlas to locate the lncRNAs within this set known to be deregulated in colon cancer, and determined which of those were also colon-specific.
The team zeroed in on a candidate called DACOR1, whose levels were lower in cancer cells than in healthy cells. Presuming this difference might have something to do with the rogue behavior of cancerous cells, Khalil asked a simple question: “What happens if we put this lncRNA back into the cancer cells?” His team re-introduced DACOR1 into its colon cancer cell lines using a lentivirus and discovered that some DNA methylation was restored, slowing the growth of cancer colonies. When the researchers mapped where DACOR1 acted on the genome after being reintroduced, they found it interacting with 150 gene loci, including 31 sites that display altered methylation in colon tumors. “[Cancer cells] deregulate the lncRNAs—and now they can change their methylation patterns to what we observe in tumors,” concludes Khalil.
“The fact that they actually demonstrate the association and how it might play a role in regulating DNA methylation is very exciting,” says Maher. “It’s not only relevant to colon cancer. I think it has broader implications.”
Interested in reading more?




